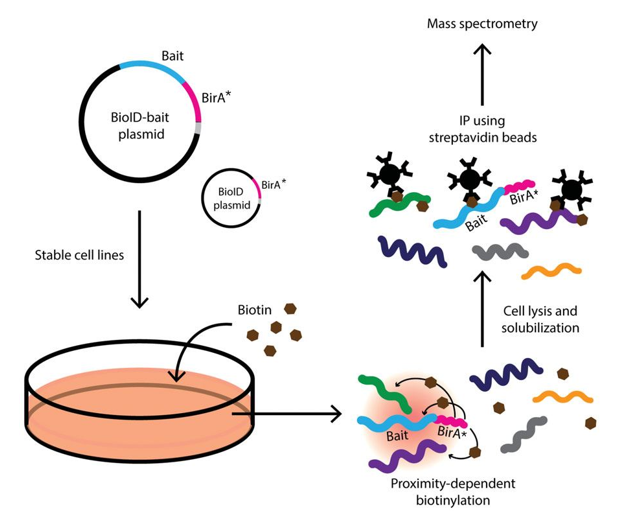
Model for application of BioID method. (a) Expression of a promiscuous... | Download Scientific Diagram
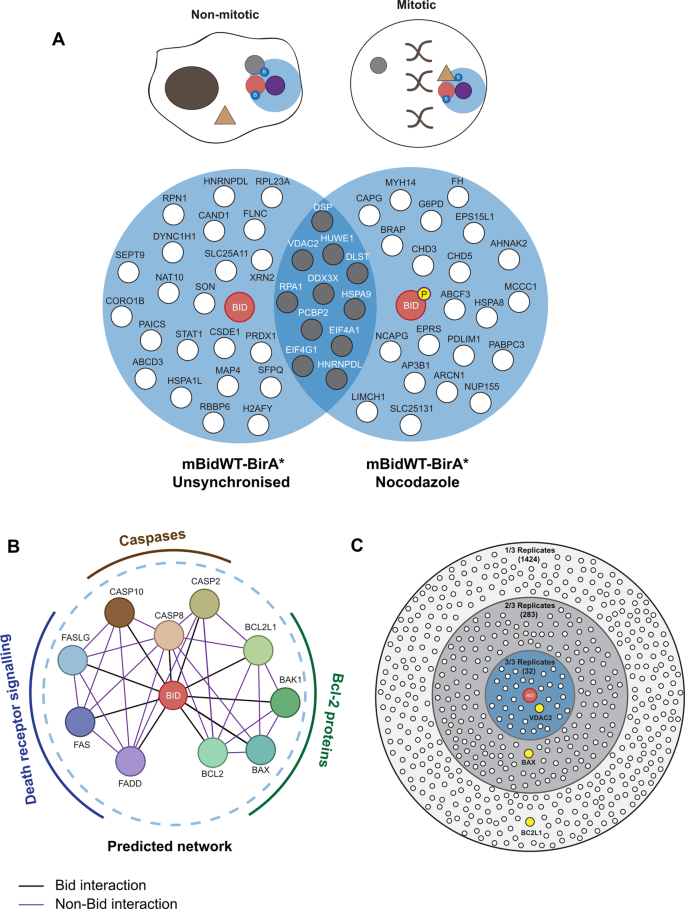
BioID-based proteomic analysis of the Bid interactome identifies novel proteins involved in cell-cycle-dependent apoptotic priming | Cell Death & Disease
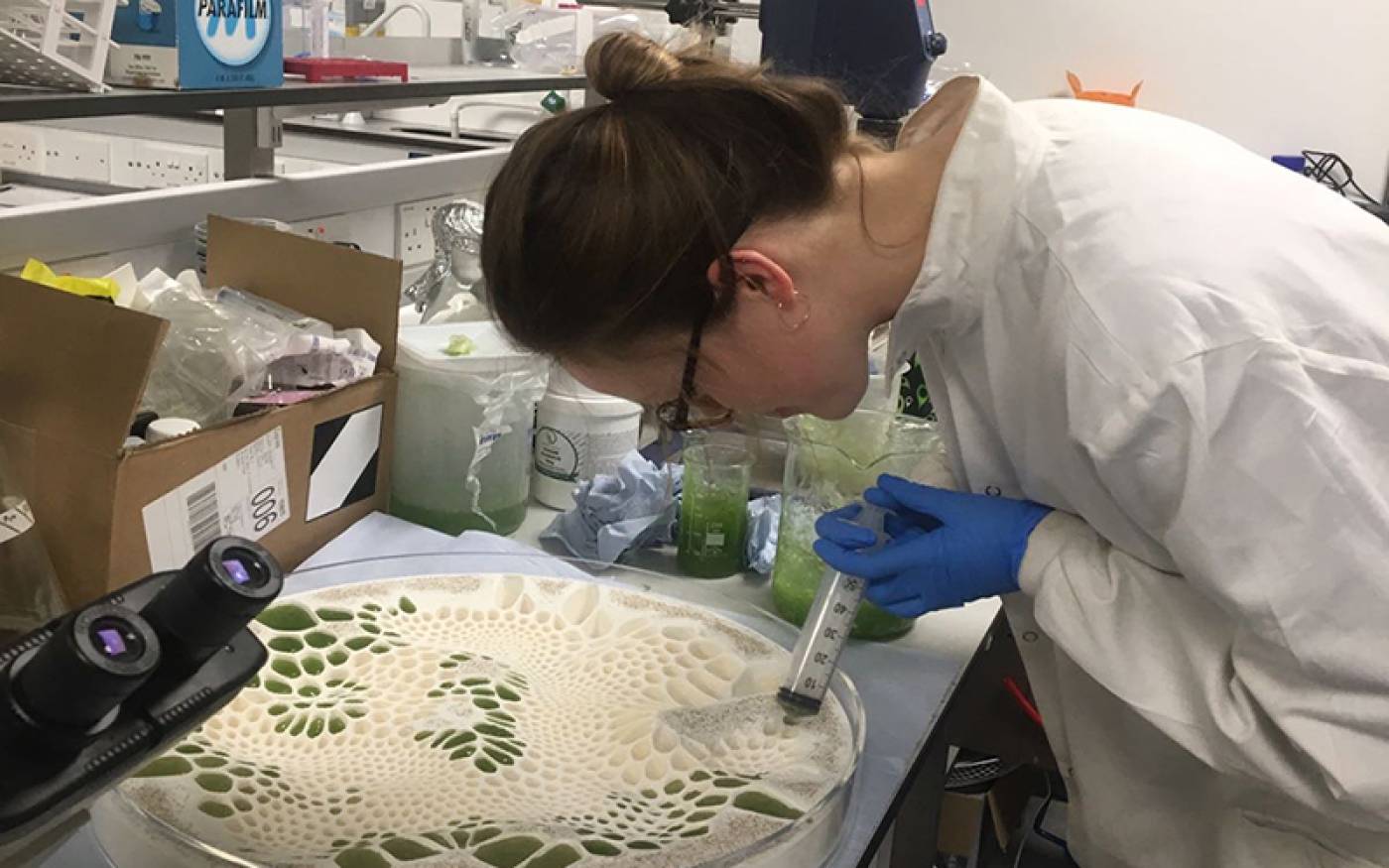

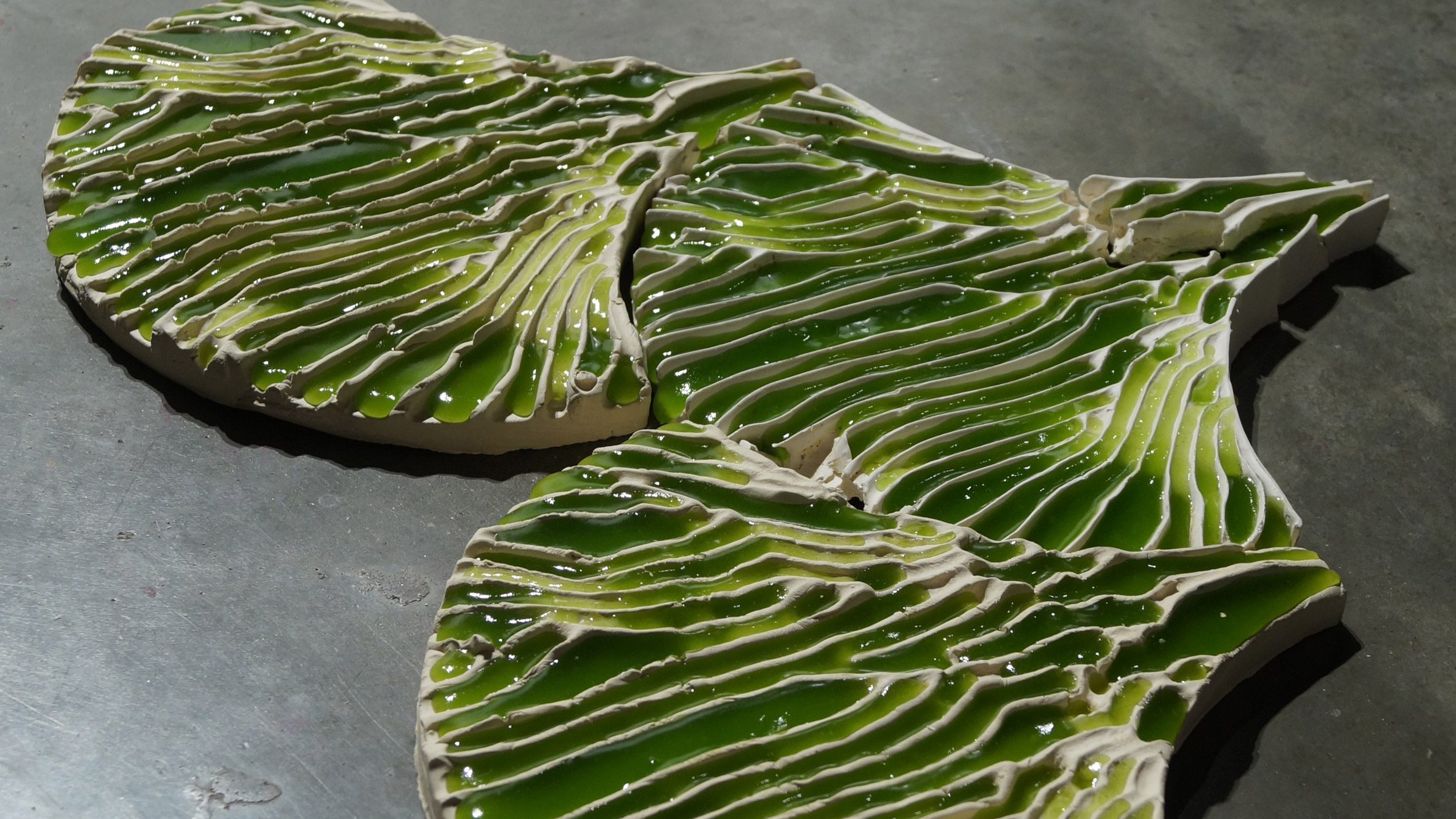
Bio-Integrated Design (Bio-ID) MArch/MSc | The Bartlett School of Architecture - UCL – University College London

In planta proximity-dependent biotin identification (BioID) identifies a TMV replication co-chaperone NbSGT1 in the vicinity of 126 kDa replicase - ScienceDirect

An AP-MS- and BioID-compatible MAC-tag enables comprehensive mapping of protein interactions and subcellular localizations | Nature Communications
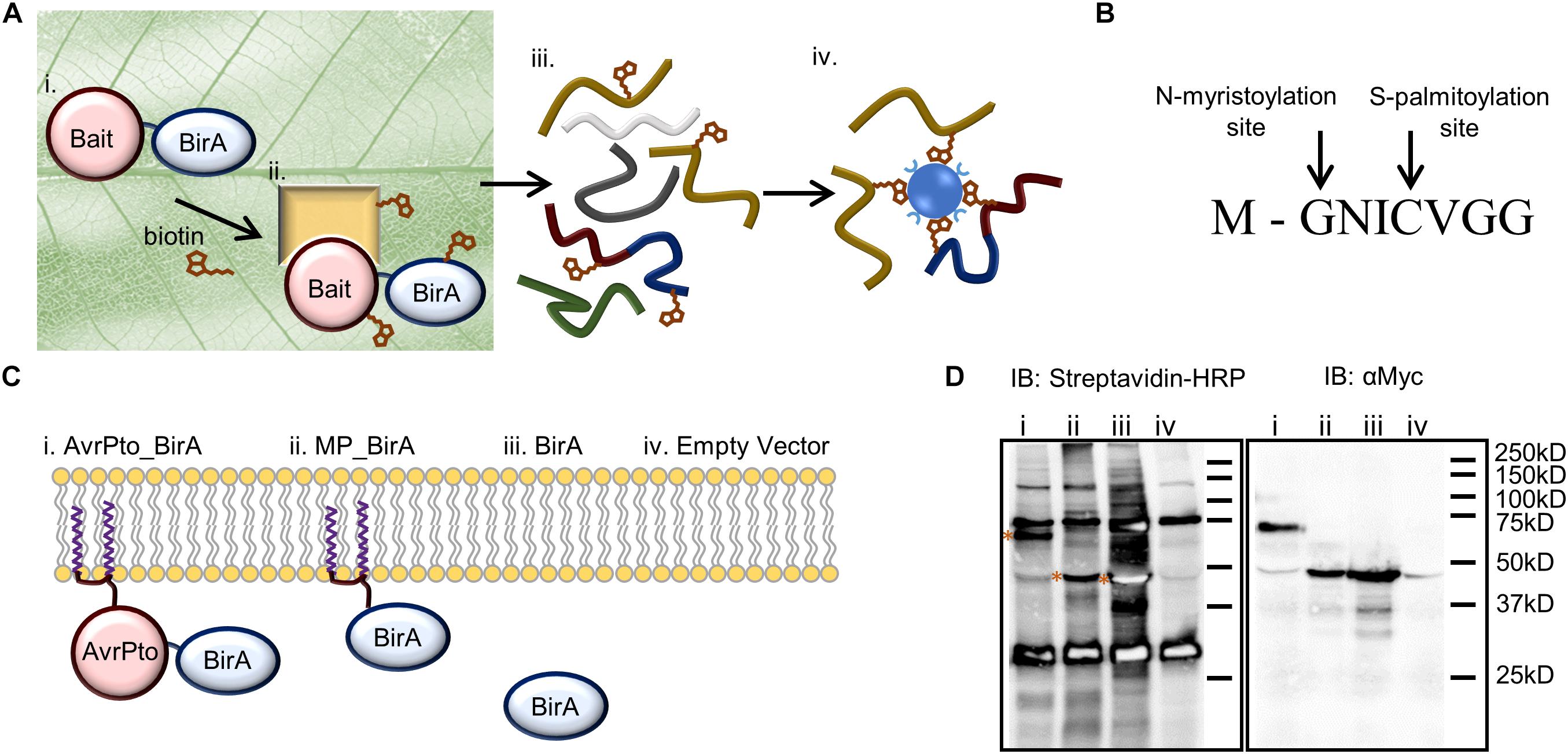
Frontiers | Development of a Rapid in planta BioID System as a Probe for Plasma Membrane-Associated Immunity Proteins
![PDF] Probing mammalian centrosome structure using BioID proximity-dependent biotinylation. | Semantic Scholar PDF] Probing mammalian centrosome structure using BioID proximity-dependent biotinylation. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/72e2710524f8b1a033ff55abc8494c4f1e1ccf5f/4-Figure1-1.png)
PDF] Probing mammalian centrosome structure using BioID proximity-dependent biotinylation. | Semantic Scholar
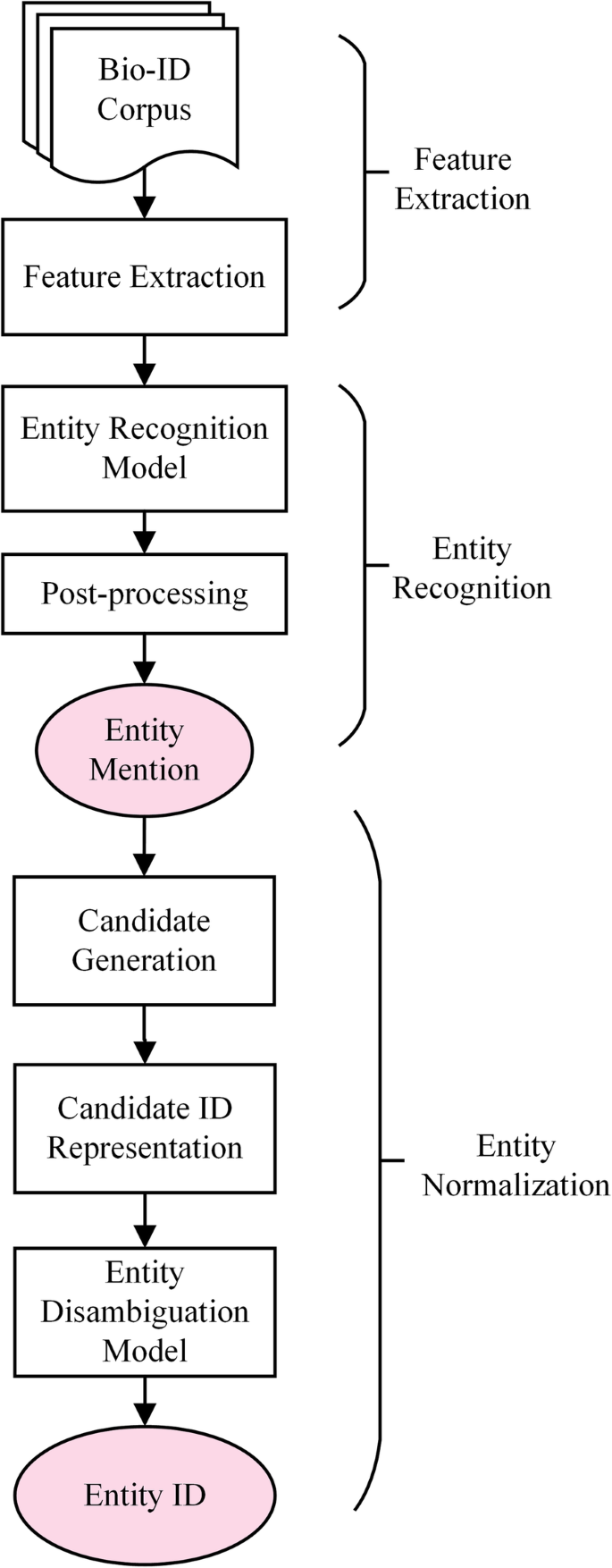
Knowledge-enhanced biomedical named entity recognition and normalization: application to proteins and genes | BMC Bioinformatics | Full Text

Proximity Labeling of Interacting Proteins: Application of BioID as a Discovery Tool - Li - 2017 - PROTEOMICS - Wiley Online Library